St Anne’s Academic Review 7 – 2017
A Modern-Day Dystopia: Can We Avoid the Post-Antibiotic Era?
Oliver Adams
STAAR 7 – October 2017, pp. 32-38
——————————–
Published: 01/10/2017
Review process: Editorial Review
It’s 2050. Long predicted, the rise of antibiotic resistance (ABR) following inadequate antibiotic stewardship, now accounts for 10 million extra deaths per year, surpassing cancer, and has inflicted some $100 trillion worth of global GDP losses. Antibiotics were once a cornerstone of modern medicine, saving countless millions of lives. They are now only sporadically effective. Most infections encountered in the clinic are caused by multi-drug (MDR) or extremely-drug (XDR) resistant bacterial strains. Many medical interventions (e.g. invasive surgery, organ transplantation, immunity-compromising chemotherapy), as well as childbirth, consequently carry a greatly increased risk of mortality. A news headline announces the death of an elderly woman following an opportunistic Klebsiella pneumoniae infection. Usually considered an innocuous commensal gut bacterium, this “superbug” was resistant to all 26 US-approved antibiotics — including those of last resort — and was thus untreatable.
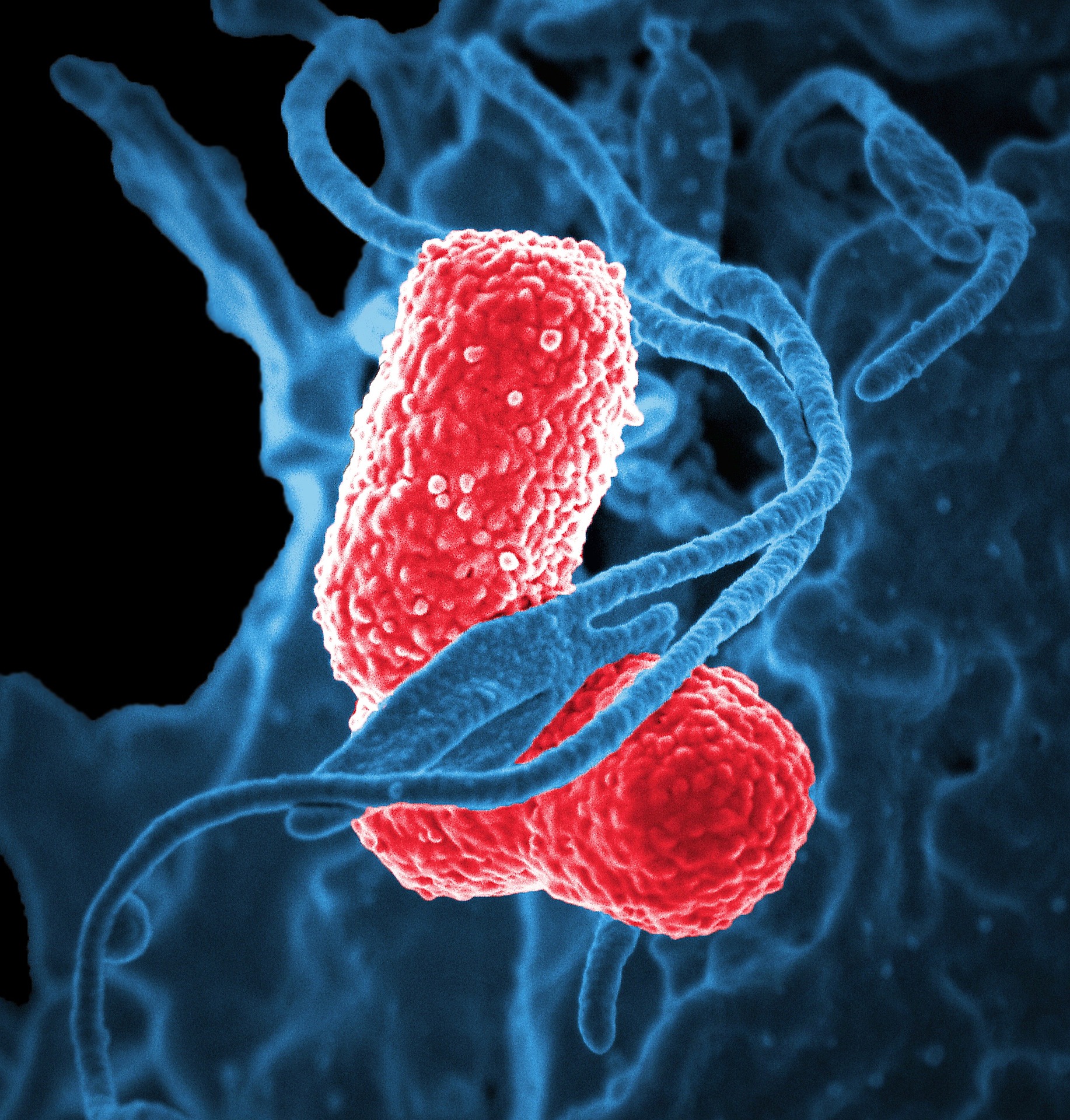 The dystopian post-antibiotic future envisaged above is constructed from predictions set out in the “Review on Antimicrobial Resistance” [1], an independent report into the implications of escalating levels of ABR, commissioned by the UK government in 2014. Unfortunately, the news report is not fictionalised. It was released in January of this year [2]. Whilst such pandrug resistance (PDR) remains thankfully rare, an epidemiological and economic crisis of global proportions continues to emerge; ABR currently estimated to account for ~700,000 deaths, and cost the US health system an excess $20 billion, per annum [3].
The dystopian post-antibiotic future envisaged above is constructed from predictions set out in the “Review on Antimicrobial Resistance” [1], an independent report into the implications of escalating levels of ABR, commissioned by the UK government in 2014. Unfortunately, the news report is not fictionalised. It was released in January of this year [2]. Whilst such pandrug resistance (PDR) remains thankfully rare, an epidemiological and economic crisis of global proportions continues to emerge; ABR currently estimated to account for ~700,000 deaths, and cost the US health system an excess $20 billion, per annum [3].
This article will start with a brief historical overview of antibiotic discovery. It will go on to look at the biochemical processes targeted by our current antibiotic armory, the multitudinous factors contributing to the increasing incidence of ABR, and the bacterial mechanisms responsible for said resistance. Finally, discussion of a range of novel approaches aimed at combatting ABR will be presented. The question is, can we avoid the post-antibiotic era?
A Historical Introduction to the Antibiotic Armory
First conceptualised as a possibility by German Immunologist and Nobel Laureate Paul Ehrlich at the beginning of the 20th century, antibiotics — or what he appealingly termed “Zauberkugel” (“Magic Bullets”) — are small molecules that possess selective bactericidal (i.e. bacterial killing) or bacteriostatic (i.e. bacterial growth inhibiting) activity without causing collateral damage to host tissues. In search of a “Magic Bullet” capable of treating sexually transmitted Treponema pallidum, the causative agent of syphilis, Ehrlich’s lab began systematically screening over 600 organoarsenic compounds for activity in infected rabbits. By 1909 they had identified Salvarsan (also known as Arsphenamine), the first successful antisyphilitic. This marked the beginning of the “antibiotic era”. Notably, variants of their pioneering high-throughput screening methodology are now standard practice in contemporary drug development pipelines [4].
The following decades saw the famous discovery (Sir Alexander Fleming, 1928), and subsequent mass production (Howard Florey & Ernst Boris Chain, 1940s), of penicillin from the fungus Penicillium notatum as well as identification of the first Sulfonamide (Prontosil); a class of synthetic bacteriostatic antibiotics derived from research into cellular dyeing compounds (Gerhard Domagk, 1930s). All four men would go on to receive the Nobel Prize in Physiology and Medicine for their achievements. Unlike Sulfonamides, but akin to penicillin, the majority of modern-day clinical antibiotics (~90%) are appropriated natural microbial products or derivatives thereof. Such molecules are part of a heterogeneous collection of specialised secreted microorganismal compounds termed “secondary metabolites”. Whilst some are known to function as bioweapons, antagonizing the growth of competing bacterial species, the ecological role of many antibiotic precursors remains to be characterized. In 1941 the term “antibiotic” was coined by microbiologist Selman Waksman, again a Nobel Laureate, to describe these inter-microbe small molecule antagonists [5].
Discovery of novel antibiotic classes — families of compounds related to one another by both chemical structure and a common antibacterial mechanism of action (see below) — then peaked in the so-called “Golden Era” between ~1940-1970. During this period the secondary metabolite profiles of readily culturable microbes (predominantly soil-dwelling bacteria of the Actinobacteria phylum) were screened for antibacterial activity by Waksman and colleagues. This natural product mining proved highly fruitful, yielding now widely prescribed antibiotics such as streptomycin (the first antitubercular agent, effective against Mycobacterium tuberculosis), tetracycline, vancomycin and chloramphenicol [4]. As the success of the “Waksman Platform” began to decline in the mid-1960s, medicinal chemistry took up the mantle. Synthetically tweaking previously isolated microbial scaffolds; chemists optimised antibacterial activity, expanded efficacy to additional pathogens, reduced toxicity and attempted to surmount the accumulating resistance to first-generation compounds. Whilst effective, the focus on pre-existing drug modification has contributed to a dearth in discovery of truly novel compounds, extending to the present-day. Despite numerous technological advances, with potential to expedite the drug development process (e.g. genome sequencing, high-throughput robotics, improvements in structural biology etc.), no new FDA (Food and Drug Administration) approved class has made it into the clinic in the past three decades [6]. This only serves to compound the ABR issue.
Mechanisms of Antibiotic Action
Antibiotics act by interfering with essential (i.e. required for microbial viability), bacterially conserved, biochemical processes and/or cellular structures. The perturbation of these triggers the associated bactericidal or bacteriostatic activity. In general, an ideal antibiotic will selectively bind and inhibit the action of one or more bacterial proteins which are absent from, or sufficiently different in, eukaryotes. This minimises off-target, side-effect inducing, activity within the patient. Despite the many hundreds of theoretically targetable aspects of bacterial physiology, our current roster of antibiotics predominantly act by only one of four mechanisms. These are: (1) compromisation of protein synthesis through inhibition of the bacterial ribosome (e.g. Aminoglycosides, Macrolides), (2) disruption of nucleic acid (i.e. DNA or RNA) associated processes (e.g. genome replication, transcription) through inhibition of RNA polymerase or DNA-maintenance enzymes (e.g. Quinolones, Ansamycins), (3) perturbation of cell wall biosynthesis, or cell membrane integrity, as to promote bacterial lysis (e.g. Beta-Lactams, Glycopeptides) and finally,(4) disruption of folate metabolism, a key molecular precursor of both DNA and amino acids (e.g. Sulfonamides, Antifolates) [4],[7]. Expanding the mechanistic repertoire of our antibiotic arsenal, as to target novel bacterial processes (e.g. fatty acid biosynthesis, quorum sensing), is an obvious goal for future drug discovery initiatives. With the above mechanisms of action in mind, ABR can now be properly explored.
Mechanisms of Resistance
ABR can be defined as the capability of a given bacterium to persist and survive at therapeutic doses of otherwise species appropriate antibiotics, rendering them useless in a clinical setting. As one may expect, given the natural origins of the majority of our antibacterial compounds, ABR is a ubiquitous biological phenomenon; having evolved over nearly 4 billion years of microbial warfare [4]. ABR therefore greatly pre-dates the comparatively recent appropriation of antibiotics by humans. It is almost certain that this ancient “resistome” — the reservoir of ABR-causative genetic determinants in the global bacterial population — features resistance mechanisms for drugs we are yet to discover [8],[9]. Multiple findings from the burgeoning field of paleomicrobiology support such dispiriting conclusions. For example, sequencing of ancient bacterial DNA preserved in Canadian permafrost identified penicillin resistance pre-dating Fleming’s discovery by some 30,000 years [10]. Comparable studies of bacteria cultured from a pristine New Mexican cave system, isolated for over 4 million years, uncovered resistance to a staggering 14 contemporary antibiotics [11]. It may be unsurprising therefore, that resistance to every licensed antibiotic has now been observed, often less than five years following initial regulatory approval. Thus the impending ABR crisis can be largely understood as the widespread human application (>70 billion clinical doses administered in 2010 [12]) , and simultaneous mismanagement (see below), of antibiotics as providing a strong selective pressure for the mobilisation of diverse resistance mechanisms which have accumulated on an evolutionary timescale.
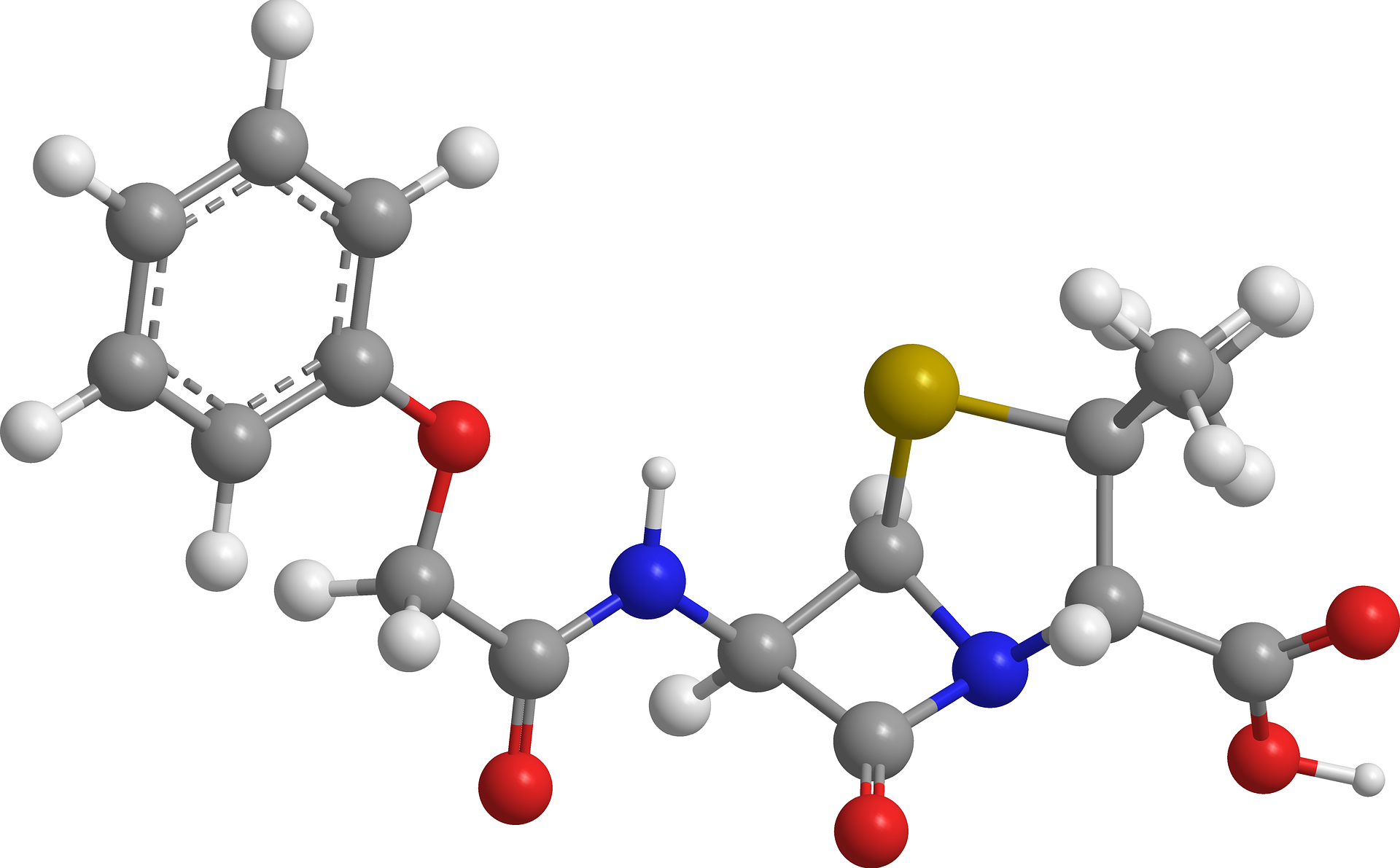
At the molecular level, ABR is achieved by at least one of three principal mechanisms, these being: (1) reduction of intracellular antibiotic concentration to below therapeutic levels, (2) modification and/or protection of the antibiotic target protein, and (3) direct enzymatic modification and/or degradation of the antibiotic. In the first instance, bacteria enact measures to reduce antibiotic permeability (e.g. down-regulating outer-membrane porins — substrate non-specific protein channels) or actively remove intracellular antibiotic through overexpression of cell surface efflux pumps. The latter proteins are often competent at transporting more than one class of antibacterials, thus promoting MDR. In the second case a point-mutation (a single amino acid substitution) or post-translational modification (an enzyme-catalysed chemical alteration) of an antibiotic target protein induces a structural change; this prevents high-affinity association with the antibiotic without detrimentally compromising the protein’s endogenous function. Alternatively, a target protein may simply be overexpressed as to prevent full inhibition on antibiotic exposure. In the final situation, bacterial enzymes catalyse the hydrolytic breakdown (e.g. extended spectrum Beta- lactamases), or inactivating chemical modification (e.g. Chloramphenicol Acetyltransferase), of the offending drug. Many thousands of such enzymes have been identified to date, often being seen to evolve altered substrate spectra to keep pace with newly synthesised antibiotic derivatives [13].
Whilst some above ABR mechanisms can be triggered by mutations within the bacterial chromosome, many are encoded on mobile genetic elements (e.g. plasmids, integrons etc.). These are capable of being exchanged between, often unrelated, bacteria by the processes of horizontal gene transfer (HGT, i.e. conjugation, transformation or transduction) and thus ABR can be rapidly spread throughout a bacterial community assuming sufficient selective pressure (i.e. antibiotic presence). Worryingly, HGT may also facilitate the accumulation of multiple resistances in a single organism; as seen in MDR, XDR and PDR bacterial strains.
Causes of Rising Antibiotic Resistance Incidence
As alluded, the increasing spread of ABR has arisen from a combination of prolonged antibiotic overuse alongside antibiotic misuse. However, humanity has long been warned about perils of ABR. Notably, clearly aware of the issue over 70 years ago, Fleming himself said:
“The thoughtless person playing with penicillin treatment is morally responsible for the death of the man who succumbs to infection with penicillin-resistant organism. I hope this evil can be averted [14].”
In addition to sizeable human antibiotic dispensation (>3 million kilograms in the US in 2009), a key issue remains the mass prophylactic and growth-promotional (i.e. non-therapeutic) application of antibiotics within agricultural industries (i.e. domestic livestock, aquaculture, apiculture etc.). The farming industry accounted for 80% of all antibiotic consumption in the US in 2010 (>13 million kilograms). Many of these drugs are concerningly also used in conventional medicine; risking expedited flow of agriculturally selected ABR into human pathogens via HGT [8].
In terms of therapeutic antibiotic use, ABR has been exacerbated by both inadequate regulation and reprehensible prescribing practices. For example, approximately a fifth of all antibiotic use outside of the US and EU operates on a non-prescription “direct-to-consumer” model. This lacks all the hallmark attributes of successful antimicrobial therapy; promoting improper drug choice, under-dosing and poor regime compliance. Even in prescription-based systems, physicians are often pressured by patients into inappropriately administering antibiotics when they are evidently not required (e.g. for viral infections) [15]. Up to ~30% of US prescribed antibiotics are thought to fall into this category [16]. Noted previously, the effects of ABR are being compounded by a stagnation in the release of new antibiotics (e.g. 16 antibacterials were FDA approved between 1983-1987, only 2 between 2008-2012). In recent years, the pharmaceutical industry has redirected investment into more profitable areas; the limited treatment duration, low cost per dose, ABR-influenced market longevity and stewardship practices (e.g. restricted prescription of new compounds) associated with antibiotics making novel antibacterial development an unattractive prospect [15].
Avoiding a Medical Dystopia
If humanity is to continue to retain the immense medical benefits bestowed by antibiotics over the past century, a concerted and sustained global approach will undoubtedly be required. This must involve interdisciplinary cooperation between the pharmaceutical and agricultural industries, the research community (e.g. microbiologists, medicinal chemists, epidemiologists), policy makers, NGOs, healthcare professionals, and educationalists; to list but a few [3]. It is beyond the scope of this piece to consider in detail all ongoing initiatives to combat ABR, but broadly, solutions fall into three main categories: (1) efforts to extend the lifespan of our current antimicrobials (i.e. antibiotic stewardship), (2) development of new antibiotics and/or therapeutic alternatives, and (3) methods to reduce the incidence of bacterial infection (i.e. reducing the need for antibacterial intervention in the first place).
Key to extending the functional lifespan of already licensed antibiotics is a move towards a system of reduced misuse and minimised consumption; ideally eliminating the non-medicinal application of antibacterials in agriculture. Whilst the EU banned growth-promotional antibiotic use in livestock in 2006, the US only recently followed suit through the FDA’s Veterinary Feed Directive in January 2017. Globally the practice remains rife [15]. Readily complementing such a worldwide ban would be implementation of systems to sever other routes responsible for environmental anthropogenic antibiotic contamination; the breakdown of antibiotic compounds in sewage, for example. By reducing the environmental antibiotic load, we should in theory lessen the selection for ABR determinants in natural bacterial ecosystems and thus the potential for ABR transfer to human pathogens. Likewise, essential to maintaining our current antibacterial roster is an improvement in prescribing practices; ensuring antibiotics are only administered when required and that an effective antibiotic is always prescribed first time. Both could be achieved through development of rapid-ABR diagnostics. Currently, the ABR-profile of a patient’s infection is determined using culture-based techniques (i.e. growing clinical isolates in the presence of various antibiotics at multiple concentrations); requiring days-to-months to complete depending on microbe growth rate. As a result, physicians are pushed to empirically prescribe broad-spectrum antibiotics before the ABR-profile is known; risking initial treatment failure. If, however, the ABR-profile could be determined in significantly less than 24 hours the physician would be empowered to both withhold antibiotics from patients with inappropriate infections as well as pick an effective drug first time [4],[8]. Although such diagnostics remain far from widespread clinical release, a frontrunner is the use of bacterial whole-genome sequencing (WGS) to directly identify the genetic determinants of ABR in a patient’s sample [17]. A successful example of this is the WGS approach recently adopted by Public Health England reference laboratories for Mycobacterial infections (e.g. tuberculosis, leprosy) [18].
 Even with improvements in antibiotic management, discovery and licensing, new antibiotics — preferably exploiting novel biochemical mechanisms — will be vital to combat resistance. Microbial evolution will always supply new ABR mechanisms to outmanoeuvre our armory. Reminiscent of the “Golden Era” of antibiotic discovery, an effective strategy would be to explore the secondary metabolite profiles of bacterial populations in previously untapped ecological niches (e.g. marine microbes). Contemporary technologies (e.g. the iChip) are also allowing screening of soil bacteria previously unculturable during Waksman’s time, an approach having yielded Teixobactin — the first antibiotic discovered with a novel mechanism of action for over three decades. It is still in early development [19]. To ensure new compounds continue to be found it is clearly essential that governments and funding bodies make efforts to re-incentivise antibiotic development. A good example is the FDA’s GAIN (Generating Antibiotic Incentives Now) legislation, which is expediting the regulatory approval, and extending the patents, of novel antimicrobials in the US. Similar incentives must be applied on an international scale [15]. Simultaneous with the search for new antibiotics are research avenues pursuing therapeutic alternatives to conventional drugs, potentially acting by mechanisms with diminished ABR-inducing potential. Such future therapies may include bacteriophage treatment (i.e. exploiting naturally antibacterial viruses for medicinal purposes), anti-virulence compounds (i.e. targeting bacterial-produced factors responsible for pathogenesis without bactericidal activity), host-targeted drugs (i.e. compounds quelling the immune response a bacterial infection, often responsible for the majority of clinical symptoms) and live biotherapeutics (e.g. introduction of bacterial populations to decolonize patients infected with MDR species). Unfortunately, the majority of these advances remain at the basic research phase. Finally, initiatives aimed at minimising the incidence of bacterial infection will always be beneficial at maintaining antibiotic viability. Broadly, technologies improving disinfection and replacing invasive healthcare practices (e.g. alternatives to intravenous drug delivery) are likely to be effective. Vaccines against ABR-strains also hold great potential [8].
Even with improvements in antibiotic management, discovery and licensing, new antibiotics — preferably exploiting novel biochemical mechanisms — will be vital to combat resistance. Microbial evolution will always supply new ABR mechanisms to outmanoeuvre our armory. Reminiscent of the “Golden Era” of antibiotic discovery, an effective strategy would be to explore the secondary metabolite profiles of bacterial populations in previously untapped ecological niches (e.g. marine microbes). Contemporary technologies (e.g. the iChip) are also allowing screening of soil bacteria previously unculturable during Waksman’s time, an approach having yielded Teixobactin — the first antibiotic discovered with a novel mechanism of action for over three decades. It is still in early development [19]. To ensure new compounds continue to be found it is clearly essential that governments and funding bodies make efforts to re-incentivise antibiotic development. A good example is the FDA’s GAIN (Generating Antibiotic Incentives Now) legislation, which is expediting the regulatory approval, and extending the patents, of novel antimicrobials in the US. Similar incentives must be applied on an international scale [15]. Simultaneous with the search for new antibiotics are research avenues pursuing therapeutic alternatives to conventional drugs, potentially acting by mechanisms with diminished ABR-inducing potential. Such future therapies may include bacteriophage treatment (i.e. exploiting naturally antibacterial viruses for medicinal purposes), anti-virulence compounds (i.e. targeting bacterial-produced factors responsible for pathogenesis without bactericidal activity), host-targeted drugs (i.e. compounds quelling the immune response a bacterial infection, often responsible for the majority of clinical symptoms) and live biotherapeutics (e.g. introduction of bacterial populations to decolonize patients infected with MDR species). Unfortunately, the majority of these advances remain at the basic research phase. Finally, initiatives aimed at minimising the incidence of bacterial infection will always be beneficial at maintaining antibiotic viability. Broadly, technologies improving disinfection and replacing invasive healthcare practices (e.g. alternatives to intravenous drug delivery) are likely to be effective. Vaccines against ABR-strains also hold great potential [8].
In conclusion, ABR is arguably one of the greatest threats to the future of humanity alongside climate change and nuclear warfare. If we are to win the arms race against our microbial foes, it is key that we act now. We can only avoid the impending dystopia if we introduce responsible regulatory practices, unearth novel antibiotics, pioneer antibiotic-alternatives, and reduce antibiotic consumption on a global-scale.
Bibliography
[1] O’Neill, J. (2016) Tackling Drug-Resistant Infections Globally: Final Report and Recommendations: The Review on Antimicrobial Resistance
[2] Chen, L. et al. (2017) Notes From the Field: Pan-Resistant New Delhi Metallo-Beta-Lactamase-Producing Klebsiella pneumoniae – Washoe County, Nevada, 2016. MMWR Morb Mortal Wkly Rep.
[3] Sugden, R., Kelly, R., and Davies, S. (2016) Combatting Antimicrobial Resistance Globally. Nat Microbiol.
[4] Aminov, R. (2010) A Brief History of the Antibiotic Era: Lessons Learned and Challenges for the Future. Front Microbiol.
[5] Clardy, J., Fischbach, M., and Currie, C. (2010) The Natural History of Antibiotics. Curr Biol.
[6] Brown, E. and Wright, G. (2016) Antibacterial Drug Discovery in the Resistance Era. Nature.
[7] Crofts, T., Gasparrini, A. and Dantas, G. (2017) Next-generation Approaches to Understand and Combat the Antibiotic Resistome. Nat Rev Microbiol.
[8] Spellberg, B., Bartlett, G., and Gilbert, D. (2013) The Future of Antibiotics and Resistance. N Engl J Med.
[9] Olaitan, A. and Rolain, J. (2016) Ancient Resistome. Microbiol Spectr.
[10] D’Costa, V. et al. (2011) Antibiotic Resistance is Ancient. Nature.
[11] Bhullar, K. et al. (2012) Antibiotic Resistance is Prevalent in an Isolated Cave Microbiome. PloS One.
[12] Van Boeckel, T. et al. (2014) Global Antibiotic Consumption 2000 to 2010: An Analysis of National Pharmaceutical Sales Data. Lancet Infect Dis.
[13] Blair, J. et al. (2015) Molecular Mechanisms of Antibiotic Resistance. Nat Rev Microbiol.
[14] Fleming, A. (1945) Penicillin’s Finder Assays its Future. New York Times.
[15] Marston, H. et al. (2016) Antimicrobial Resistance. J Am Med Assoc.
[16] Fleming-Dutra, K. et al. (2016) Prevalence of Inappropriate Antibiotic Prescriptions Among US Ambulatory Care Visits, 2010-2011. J Am Med Assoc.
[17] Didelot, X. et al. (2012) Transforming Clinical Microbiology with Bacterial Genome Sequencing. Nat Rev Genet.
[18] Walker, T. et al. (2017) Tuberculosis is Changing. Lancet Infect Dis.
[19] Ling, L. et al. (2015) A New Antibiotic Kills Pathogens Without Detectable Resistance. Nature.

A Modern-Day Dystopia: Can We Avoid the Post-Antibiotic Era? by Oliver Adams is licensed under a Creative Commons Attribution 4.0 International License.
<< Back to Contents
<< Back to Publications
St Anne's Academic Review (STAAR) A Publication by St Anne's College Middle Common Room ISSN 2048-2566 (Online) ISSN 2515-6527 (Print)

